Speaker: Markus Töpfer (VRVis)
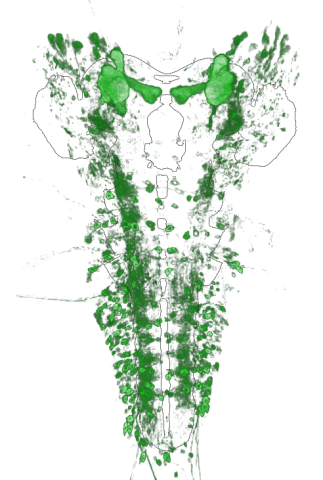
The integration of modern genetic techniques, advanced volumetric imaging methods, and single-cell extraction methods has empowered neurobiologists to create extensive digitized and coregistered specimen sample collections. These collections serve as valuable resources for studying neuronal structures, functional compartments, and neurological connections within the brain. By sampling single cells from various locations within the animal brain, scientists can investigate cell type distributions and gene expressions. However, the exploration of these vast collections, which include volumetric images, segmented structures, and gene expression data, poses a significant challenge in neuroscience. Efficient access to specific regions of interest in all images, derivative processed data, cell samples, and metadata is essential for researchers. In this talk, I present a flexible and extensible approach to spatially index and store volumetric grid data and region-based data, enabling efficient access and providing a streamlined method for implementing new data abstractions, query types, and indexing strategies. The framework supports different datasets, the encoding of neurological structural types, and incorporates a layering mechanism to handle multiple data representations or time-dependent data within a single data structure. Standardized interfaces are defined for loading voxel and region data, preprocessing them, creating data abstractions, and implementing new query types. The data storage is managed using a storage engine approach, allowing users to leverage different storage mechanisms or introduce their own.